Draws plausible values based on test scores
Arguments
- dataSrc
a connection to a dexter database, a matrix, or a data.frame with columns: person_id, item_id, item_score
- parms
An object returned by function
fit_enormcontaining parameter estimates or a data.frame with columns item_id, item_score and, beta. If parms are provided, item parameters are considered known, see @details. If parms is NULL, they will be estimated Bayesianly.- predicate
an expression to filter data. If missing, the function will use all data in dataSrc
- covariates
name or a vector of names of the variables to group the populations used to improve the prior. A covariate must be a discrete person property (e.g. not a float) that indicates nominal categories, e.g. gender or school. If dataSrc is a data.frame, it must contain the covariate.
- nPV
Number of plausible values to draw per person.
- parms_draw
when the item parameters are estimated with method "Bayes" (see:
fit_enorm), parms_draw specifies whether to use a sample (a different item parameter draw for each plausible values draw) or the posterior mean of the item draws. Alternatively, it can be an integer specifying a specific draw. It is ignored when parms is not estimated Bayesianly.- link_error
only for cml, whether to use the maximum likelihood estimates or random draws based on the covariance matrix, thus including estimation/linking error in the pv draws.
- prior_dist
use a normal prior for the plausible values or a mixture of two normals. A mixture is only possible when there are no covariates.
- merge_within_persons
If a person took multiple booklets, this indicates whether plausible values are generated per person (TRUE) or per booklet (FALSE)
Value
A data.frame with columns booklet_id, person_id, booklet_score, any covariate columns, and nPV plausible values named PV1...PVn.
Details
When the item parameters are estimated using fit_enorm(..., method='Bayes') and parms_draw = 'sample',
or when they are estimated with CML and lin_error=TRUE the uncertainty
of the item parameter estimates is taken into account when drawing multiple plausible values. To use these options, parms must be an object of type enorm and not a data.frame.
In there are covariates, the prior distribution is a hierarchical normal with equal variances across groups. When there is only one group this becomes a regular normal distribution. When there are no covariates and prior_dist = "mixture", the prior is a mixture distribution of two normal distributions which gives a little more flexibility than a normal prior.
References
Marsman, M., Maris, G., Bechger, T. M., and Glas, C.A.C. (2016) What can we learn from plausible values? Psychometrika. 2016; 81: 274-289. See also the vignette.
Examples
db = start_new_project(verbAggrRules, ":memory:",
person_properties=list(gender="<unknown>"))
add_booklet(db, verbAggrData, "agg")
#> no column `person_id` provided, automatically generating unique person id's
#> $items
#> [1] "S1DoCurse" "S1DoScold" "S1DoShout" "S1WantCurse" "S1WantScold"
#> [6] "S1WantShout" "S2DoCurse" "S2DoScold" "S2DoShout" "S2WantCurse"
#> [11] "S2WantScold" "S2WantShout" "S3DoCurse" "S3DoScold" "S3DoShout"
#> [16] "S3WantCurse" "S3WantScold" "S3WantShout" "S4DoCurse" "S4DoScold"
#> [21] "S4DoShout" "S4WantCurse" "S4WantScold" "S4WantShout"
#>
#> $person_properties
#> [1] "gender"
#>
#> $columns_ignored
#> [1] "anger"
#>
add_item_properties(db, verbAggrProperties)
#> 4 item properties for 24 items added or updated.
f=fit_enorm(db)
pv_M=plausible_values(db,f,(mode=="Do")&(gender=="Male"))
pv_F=plausible_values(db,f,(mode=="Do")&(gender=="Female"))
oldpar = par(mfrow=c(1,2))
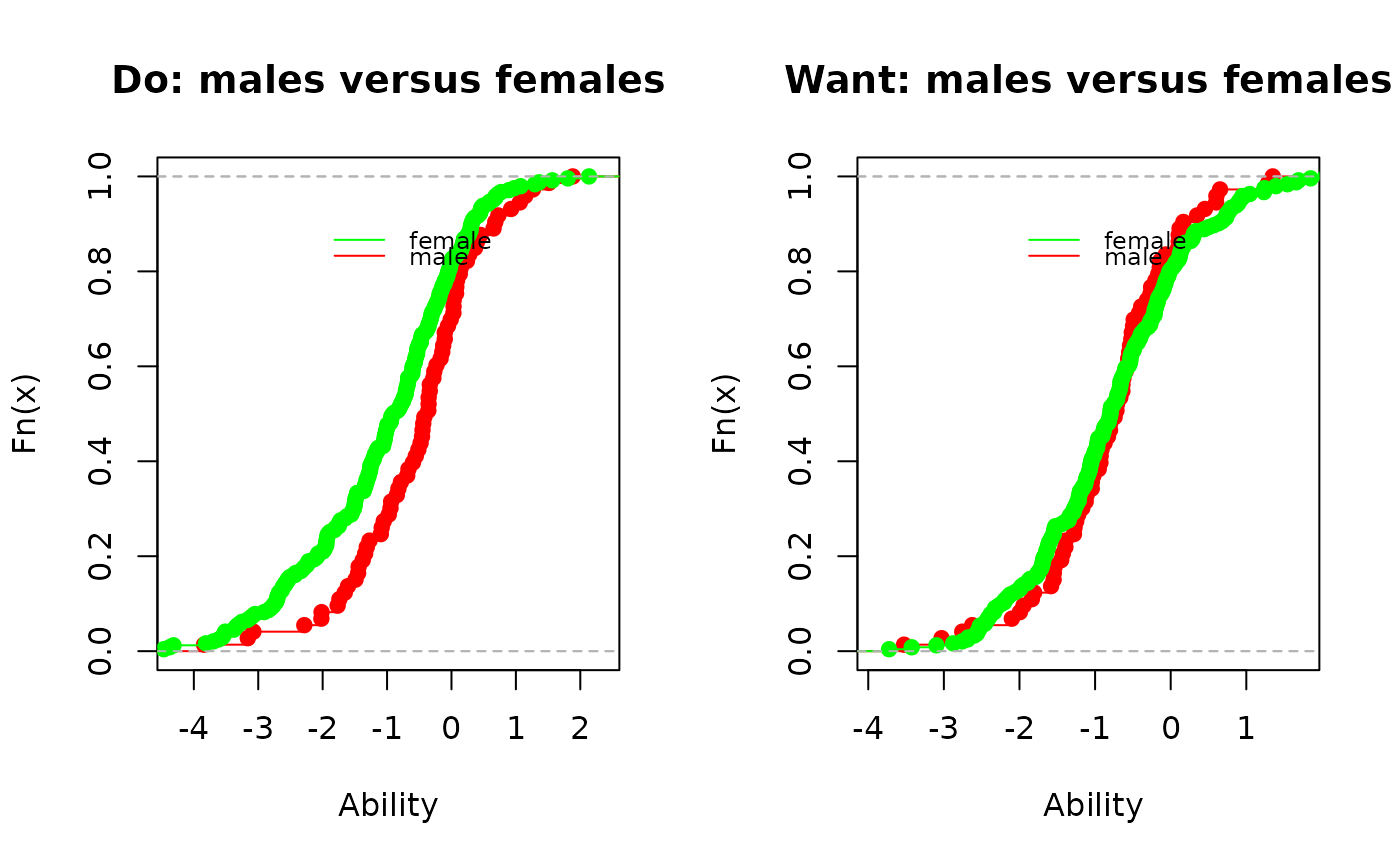
plot(ecdf(pv_M$PV1),
main="Do: males versus females", xlab="Ability", col="red")
lines(ecdf(pv_F$PV1), col="green")
legend(-2.2,0.9, c("female", "male") ,
lty=1, col=c('green', 'red'), bty='n', cex=.75)
pv_M=plausible_values(db,f,(mode=="Want")&(gender=="Male"))
pv_F=plausible_values(db,f,(mode=="Want")&(gender=="Female"))
plot(ecdf(pv_M$PV1),
main="Want: males versus females", xlab=" Ability", col="red")
lines(ecdf(pv_F$PV1),col="green")
legend(-2.2,0.9, c("female", "male") ,
lty=1, col=c('green', 'red'), bty='n', cex=.75)
 par(oldpar)
close_project(db)
par(oldpar)
close_project(db)